Pac-Man-like CRISPR enzymes have potential for disease diagnostics
Researchers find 10 new kinds of Cas13a enzyme which, when activated, indiscriminately chew up all RNA, like in the old Pac-Man arcade game
May 4, 2017
University of California, Berkeley, researchers have described 10 new CRISPR enzymes that, once activated, behave like Pac-Man to chew up RNA in a way that could be used as sensitive detectors of infectious viruses.

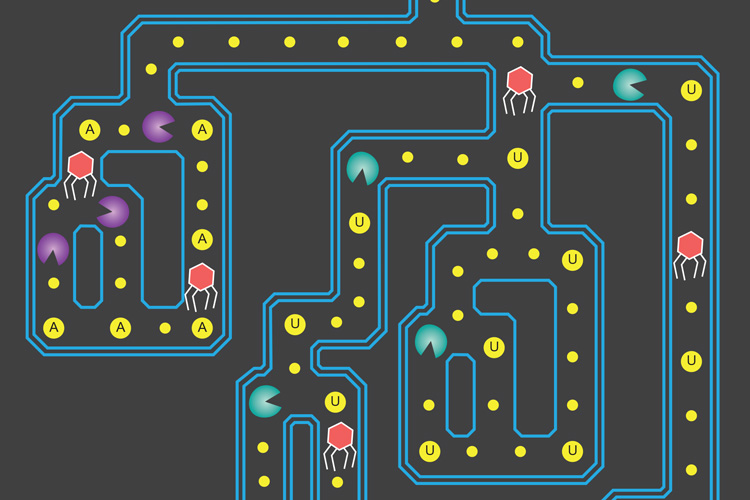
CRISPR-Cas13a enzymes act like the 1980s arcade game Pac-Man, where one type (purple) chomps on RNA at the nucleotide adenine (A), while another type (bluegreen) cuts RNA at the nucleotide uracil (U). The red spider-like figures represent viruses that attack bacteria, the real-life target of these CRISPR enzymes. Alexandra East-Seletsky graphic.
The new CRISPR enzymes are variants of a CRISPR protein, Cas13a, which the UC Berkeley researchers reported last September in Nature could be used to detect specific sequences of RNA, such as from a virus. They showed that once CRISPR-Cas13a binds to its target RNA, it begins to indiscriminately cut up all RNA, easily cutting RNA linked to a reporter molecule, making it fluoresce to allow signal detection.
Two teams of researchers at the Broad Institute subsequently paired CRISPR-Cas13a with RNA amplification, and showed that the system, which they dubbed SHERLOCK, could detect viral RNA at extremely low concentrations, detecting the presence of dengue and Zika viral RNA, for example.
Such a system could be used to detect any type of RNA, including RNA distinctive of cancer cells.
While the original Cas13a enzyme used by the UC Berkeley and Broad teams cuts RNA at one specific nucleic acid, uracil, three of the new Cas13a variants cut RNA at adenine. This difference allows simultaneous detection of two different RNA molecules, such as from two different viruses.
“We have taken our foundational research a step further in finding other homologs of the Cas13a family that have different nucleotide preferences, enabling concurrent detection of different reporters with, say, a red and a green fluorescent signal, allowing a multiplexed enzymatic detection system,” said first author Alexandra East-Seletsky, a UC Berkeley graduate student in the laboratory of Jennifer Doudna, one of the inventors of the CRISPR-Cas9 gene-editing tool. East-Seletsky was also a co-first author of the September Nature paper.
East-Seletsky, Doudna and their UC Berkeley colleagues reported their findings May 4 in the journal Molecular Cell.
RNA killing spree
The CRISPR-Cas13a family, formerly referred to as CRISPR-C2c2, is related to CRISPR-Cas9, which is already revolutionizing biomedical research and treatment because of the ease of targeting it to unique DNA sequences to cut or edit. While the Cas9 protein cuts double-stranded DNA at specific sequences, the Cas13a protein – a nucleic acid-cutting enzyme referred to as a nuclease – latches onto specific RNA sequences, and not only cuts that specific RNA, but runs amok to cut and destroy all RNA present.
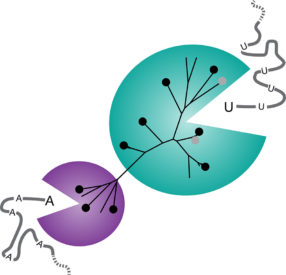
Berkeley researchers tested 10 newly discovered Cas13a enzymes and found that they fall into two groups: a larger group (bluegreen) that cuts RNA at the nucleotide uracil (U) and a smaller group (purple) that cuts at the nucleotide adenine (A). The black balls represent homologs of Cas13a that were active; the grey balls homologs that were inactive. Alexandra East-Seletsky graphic.
“Think of binding between Cas13a and its RNA target as an on-off switch — target binding turns on the enzyme to go be a Pac-Man in the cell, chewing up all RNA nearby,” East-Seletsky said. This RNA killing spree can kill the cell.
In their September Nature paper, the UC Berkeley researchers argued that the Pac-Man activity of CRISPR-Cas13a is its main role in bacteria, aimed at killing infectious viruses or phages. As part of the immune system of some bacteria, it allows infected cells to commit suicide to save their sister microbes from infection. Similar non-CRISPR suicide systems exist in other bacteria.
The UC Berkeley researchers subsequently searched databases of bacterial genomes and found 10 other Cas13a-like proteins, which they synthesized and studied to assess their ability to find and cut RNA. Of those, seven resembled the original Cas13a, while three differed in where they cut RNA. RNA, which serves many functions inside the cell, including as messenger RNA – working copies of DNA – consists of four different nucleotides: adenine, cytosine, guanine and uracil.
“Building on our original work, we now show that it is possible to multiplex these enzymes together, extending the scope of the technology,” East-Seletsky said. “There is so much diversity within the CRISPR-Cas13a family that can be utilized for many applications, including RNA detection.”
Doudna, a professor of molecular biology and of chemistry and a Howard Hughes Medical Institute investigator, noted that detection of infectious RNA may or may not require amplification, which is a complicated step.
“Our intention is to develop the Cas13a family of enzymes for point-of-care diagnostics that are robust and simple to deploy”, Doudna said.
Co-authors with East-Seletsky and Doudna are former UC Berkeley postdoctoral fellow Mitchell O’Connell, now an assistant professor at the University of Rochester, and UC Berkeley postdocs David Burstein and Gavin Knott. The work was supported in part by a Frontiers Science award from the Paul Allen Institute and by the National Science Foundation (MCB-1244557).
RELATED INFORMATION