New drug strategy: Target ribosome to halt protein production
LDL-lowering drug selectively stalls ribosome, suggesting other potential drugs that target ribosome
March 21, 2017
The discovery of a chemical compound that halts the production of a small set of proteins while leaving general protein production untouched suggests a new drug search strategy: Find compounds that target undesired proteins before they are even made.
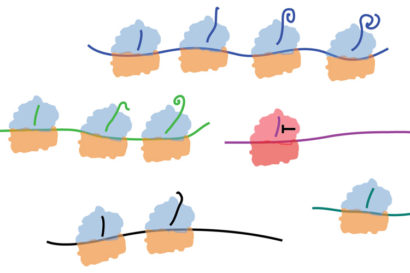
Ribosomes lined up along pieces of messenger RNA extrude proteins that curl up once they emerge from the ribosome’s internal tunnel. UC Berkeley and Pfizer scientists discovered that a small molecule (black T) can kink the growing protein inside the tunnel and stall its production while leaving other protein production unaffected. Jamie Cate image.
Many of today’s therapies for cancer or heart disease are monoclonal antibodies that bind and disable proteins outside the cell. The immunotherapeutic checkpoint inhibitors, such as Yervoy, block suppressor proteins, for example, unleashing the immune system to attack cancer.
But monoclonal antibodies aren’t effective against all proteins, can’t enter cells and must be delivered via injection.
In a paper appearing today in the journal PLOS Biology, researchers at the University of California, Berkeley, and Pfizer Worldwide Research and Development report finding a small molecule that was able to block the production of a specific protein involved in LDL (low-density lipoprotein) turnover by stalling only the ribosome that produces that protein. Ribosomes are large, general-purpose molecular machines that translate genetic instructions in the form of messenger RNA into the proteins used to build cells, the enzymes in charge of cellular housekeeping, and the hormones that carry messages in and between cells.
When delivered orally to rats, the small molecule lowered LDL cholesterol levels, much the way statins do, though by a different mechanism: by lowering the production of the protein PCSK9.
While antibiotics like erythromycin are known to stall the ribosome, they halt production of most proteins, said Jamie Cate, one of two senior authors, a UC Berkeley professor of molecular and cell biology and of chemistry and a faculty scientist at Lawrence Berkeley National Laboratory.
The chemical in this instance stalls the ribosome only when it’s producing the protein PCSK9 and a couple of dozen others out of the tens of thousands of proteins the body produces, as shown by a relatively new technique called ribosomal profiling.
“PCSK9 was just where we started. Now we can think about how to come up with other small molecules that hit proteins that nobody has been able to target before because, maybe, they have a floppy part, or they don’t have a nook or cranny where you can bind a small molecule to inhibit them,” Cate said. “This research is saying, we may be able to just prevent the synthesis of the protein in the first place.”
Cate suspects that the small molecule in the current study, a multi-ringed chlorinated compound, could serve as a template, like a key blank that can be machined to open a specific lock.
“We now have this key blank that we can cut in a number of different ways to try to go after undruggable proteins in a number of different disease states,” Cate said. “No one really thought that would have been possible before.”
Stalling the ribosome
The small molecule was discovered by Pfizer labs through live-cell screening for compounds that lower production of the protein PCSK9 (proprotein convertase subtilisin kexin 9), which regulates the recycling of the LDL receptor. Knocking out the protein is known to lower blood levels of LDL cholesterol, the so-called bad cholesterol, presumably lowering risk of cardiovascular disease. PCSK9 inhibitors, mostly monoclonal antibodies, actually lower LDL better than the well-known statins, though they have to be injected into the bloodstream.
When it became clear that the chemical was acting on the ribosome, Spiros Liras, vice president of medicinal chemistry at Pfizer, approached Cate and Jennifer Doudna, both leaders in the field of ribosome function and translation, to establish a collaboration through UC Berkeley’s California Institute for Quantitative Bioscience (QB3) to further investigate the questions of selectivity and mechanism of action. Cate is also director of UC Berkeley’s Center for RNA Systems Biology, while Doudna is a professor of molecular and cell biology and of chemistry, a Howard Hughes Medical Institute investigator and executive director of the Innovative Genomics Institute.
“Pfizer brought a significant depth of knowledge and resources to the collaboration, including fundamental cell biology, disease-relevant expertise, chemical biology and medicinal chemistry,“said Liras. “We aimed at building a strong cross-institutional collaboration which would complement our strengths in drug discovery with UC Berkeley’s strengths in ribosome biochemistry and structural biology.“
In the PLOS Biology paper, Cate, Robert Dullea at Pfizer and their teams at UC Berkeley and Pfizer describe how the drug interacts with the ribosome to halt protein production.
According to Cate, the ribosome assembles amino acids into a chain inside a tunnel that holds about 30 to 40 amino acids before the end begins to poke out of the tunnel. The chemical studied appears to bind to specific amino acid sequences of the growing protein within that tunnel in the ribosome and make them kink enough to stop progress down the tunnel, halting protein synthesis.
“We found that the proteins that are stalled are too short to stick outside the ribosome,” Cate said. “So we think the compound is actually trapping this snake-like chain, the starting part of the protein, in the tunnel — not completely blocking the tunnel, but just partially blocking it, in a way that prevents this particular protein from making its way out.”
While it’s still unclear what the two dozen proteins affected have in common that makes them susceptible to stalling by the small molecule, Cate sees these findings as clear evidence that ribosomal stalling can occur very specifically, something most researchers thought unlikely.
“We think that we now have enough understanding of the mechanism that we have our foot in the door to explore the relevance of this biology more broadly,” said Cate.
Co-authors of the paper are Cate, Doudna and postdoctoral scholar Nathanael Lintner of UC Berkeley and Pfizer researchers Kim McClure, Donna Petersen, Allyn Londregan, David Piotrowski, Liuqing Wei, Jun Xiao, Michael Bolt, Paula Loria, Bruce Maguire, Kieran Geoghegan, Austin Huang, Tim Rolph and Spiros Liras.
The work was funded by Pfizer, with computing and gene sequencing assistance through resources supported by the National Institutes of Health.
RELATED INFORMATION