Scientists track frog-killing fungus to help curb its spread
Frog skin swabs reveal when and where the deadly disease has popped up in wildlife populations around the world
September 23, 2019
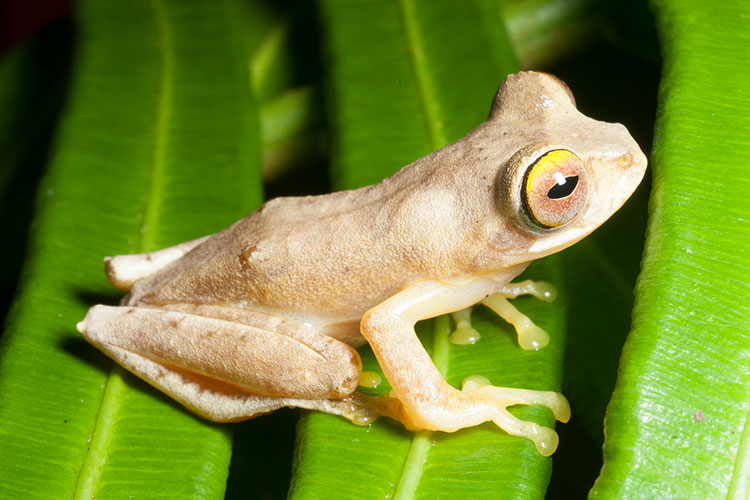

This Rhacophorus bimaculatus frog, a native of the Philippines that makes its home in the misty spray zones beneath waterfalls, is one of the many frog species imperiled by the deadly chytrid fungus. (Photo by Rafe Brown & Jason Fernandez)
From habitat loss to climate change, amphibians around the world face immense threats to their survival. One emerging and sinister threat is the chytrid fungus, a mysterious pathogen that kills amphibians by disrupting the delicate moisture balance maintained by their skin, and that is decimating frog populations around the world.
“Amphibians are already one of the most imperiled groups on the planet, and this fungal disease is further threatening their biodiversity,” said Erica Bree Rosenblum, an associate professor of environmental science, policy and management at the University of California, Berkeley.
With the help of advanced genetic testing and hundreds of frog skin swabs, Rosenblum, along with UC Berkeley graduate student Allison Byrne and an international team of collaborators, has created the most complete map to date of when and where different genetic variants of the fungus — analogous to different strains of viruses like the flu — have infected frog populations around the world.
Some of these genetic variants are deadlier than others, so knowing their current geographic distribution is key to preventing future spread of the disease, the researchers said. The investigation also uncovered a whole new genetic lineage of the fungus, one that appears to have originated in Asia and may be the oldest variant yet discovered.
“An invisible aspect of globalization is that when we move plants and animals around, we are moving their diseases around, and that can have really devastating consequences,” Rosenblum said. “If we know what lineages are where, we can better predict conservation outcomes, because some of these lineages are really deadly, and others less so.”
The study appears online the week of Sept. 23 in the journal Proceedings of the National Academy of Sciences.
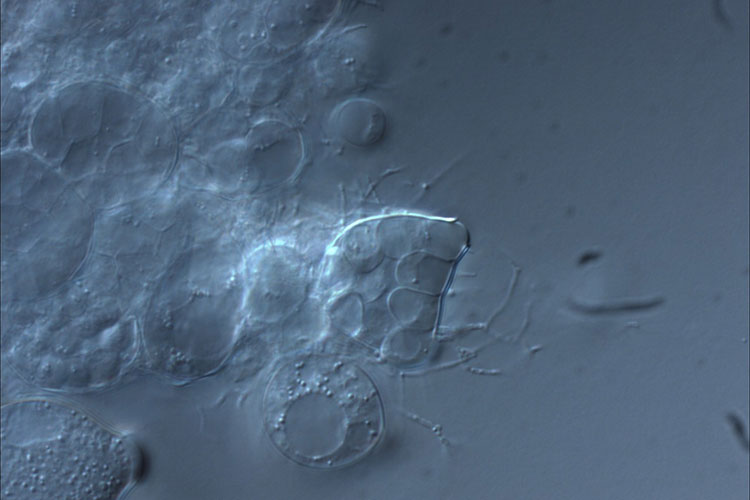
The chytrid fungus, bearing the scientific name Batrachochytrium dendrobatidis, infects frogs’ skin and disrupts the animal’s ability to properly maintain moisture and electrolyte balance. (Photo by Erica Bree Rosenblum)
“Chytrid” is the name of not one, but around 1,000 different species of fungi, most of which are harmless detritivores that spend their lives munching away on dead and rotting organic matter. The frog-killing variety, which bears the tongue-twisting name Batrachochytrium dendrobatidis, wasn’t discovered until the late 1990s, when scientists were desperately searching for the source of disturbing frog die-offs that had been popping up worldwide since the 1970s.
Though researchers now have a culprit, much remains unknown about this enigmatic disease: Some species of frog, like the American Bullfrog, are not affected by the fungus, while for others, exposure to the fungus means almost certain death.
Scientists are also not clear on the origins of the frog-killing chytrid species. It may have been lurking in some corner of the planet for thousands of years and only brought to worldwide attention when global trade of frogs for their meat and for lab experiments exposed new populations to the pathogen. On the other hand, it’s possible that a recent genetic mutation in the fungus suddenly made it more virulent, Rosenblum said.
“Not all amphibian species are equally susceptible, not all strains of the fungus are equally deadly, and there are a lot of open questions about the disease. Are humans moving it around? Are different strains coming together and mixing and becoming even more deadly?” Rosenblum said. “It’s like a murder mystery we are trying to solve.”
To find the answers, researchers often turn to genetic testing to “get under the hood” of a disease, using DNA signatures to map out different variants of a pathogen and trace them back to their source. But full genome sequencing of the chytrid fungus is notoriously difficult, requiring large samples of the fungus that must be grown and cultured in a lab and that sometimes involve killing the infected frog.
In the current study, Byrne borrowed technologies from medical research to create a new genetic test that can determine what variant of the fungus a frog has using only a skin swab. Rather than sequencing the entire genome, the test looks only at key snippets of DNA that help distinguish one variant from another. Because it only relies on small snippets, it requires much less DNA overall.
“People have been using Q-tip swabbing to test simply whether the pathogen is present or not, but now we can use the same swabs to figure out genetic details about the fungus, too,” Rosenblum said. “There are literally hundreds of thousands of these swabs sitting in people’s freezers around the world. Now, from these skin swabs, we can figure out not just that this frog is infected, but specifically what lineage of the disease it carries. Is it the same lineage that infects the frog living next door, or in another country, or across the world?”

This delicate treefrog commonly called a “Rain Frog” is only found among understory tree leaves in rainforests of the Philippines. This species of frog has also tested positive for the chytrid fungus. (Photo by Rafe Brown & Jason Fernandez)
Rosenblum and Byrne partnered with collaborators around the world to analyze 222 fungus samples collected from 24 different countries. These samples included skin swabs collected in the field, as well as some taken from preserved museum frog specimens, the oldest of which dates to 1984.
“I think something that our collaborators are most excited about is going back in time with these museum specimens,” Byrne said. “You can take an amphibian in a jar, collected a hundred years ago, swab it and now start to really build the picture of how these lineages have spread.”
The analysis revealed the distribution of the four known lineages around the globe and uncovered a whole new lineage that appears to have originated in Asia. It also showed how, in many places, different lineages of the pathogen appear to live side-by-side. This is a frightening prospect, as recent lab work has hinted that, like many human viruses, different lineages of the pathogen can mix and become deadlier.
Policies that limit the transport of frogs and other animals across international borders could help prevent the spread of the disease, the researchers say. For example, restrictions on salamander imports from Europe have so far kept a related species of the fungus, Batrachochytrium salamandrivorans, out of the United States.
“We are seeing that policy actions can slow or prevent the spread of these diseases,” Byrne said. “I think there might be a little bit of complacency around the fact that this chytrid is basically everywhere. But now we know that they are not all the same, the risks depend on what lineage is there, and then when you get this mixing of lineages, you could have many unintended consequences.”
Co-authors of the paper include Vance T. Vredenburg, Raul Figueroa-Valenzuela and Hasan Sulaeman of San Francisco State University; An Martel and Frank Pasmans of Ghent University; Rayna C. Bell of the California Academy of Sciences; David C. Blackburn of the Florida Museum of Natural History; Cheryl J. Briggs of UC Santa Barbara; Rafe M. Brown of the University of Kansas; Molly C. Bletz of the University of Massachusetts, Boston; Jaime Bosch of the Museo Nacional de Ciencias Naturales; Alessandro Catenazzi of Florida International University; Mariel Familiar Lopez of Griffith University; Sonia L. Ghose of UC Davis; Jef R. Jaeger and Anthony W. Waddle of the University of Nevada, Las Vegas; Andrea J. Jani of the University of Hawaii at Manoa; Miloslav Jirku of the Czech Academy of Sciences; Roland A. Knapp of the Sierra Nevada Aquatic Research Laboratory; Antonio Munoz of El Colegio de la Frontera Sur, San Cristobal de las Casas, Mexico; Daniel M. Portik of the University of Arizona, Tuscon; Corinne L. Richards-Zawacki of the University of Pittsburgh; Heidi Rockney of Oregon State University; Sean Rovito of Reptile, Amphibian and Fish Conservation Netherlands; Tariq Stark of Centro de Investigacion y de Estudios Avanzados del Instituto Politecnico Nacional, Irapuato, Mexico; Nguyen Thien Tao of the Vietnam Academy of Science and Technology; Jamie Voyles of the University of Nevada, Reno; and Zhiyong Yuan of the Southwest Forestry University, China.
This work was supported by the National Science Foundation (IOS-1354241, DEB-1557190, DEB-1551488, DEB-1202609, DEB-0743491, 0334952, 1418895, 1654388, 1633948 and an NSF GRFP), the National Institutes of Health (NIH COBRE Phase III grant P30GM103324), the Czech Science Foundation and Institute of Parasitology (P506/10/2330 and RVO: 60077344), the National Natural Science Foundation of China (31702008) and the Applied Basic Research Project of Yunnan (2018FD047).
RELATED INFORMATION